SAX-7/L1CAM and HMR-1/cadherin function redundantly in blastomere compaction and non-muscle myosin accumulation during Caenorhabditis elegans gastrulation.
Theresa M. Grana1, Cox, E.A.2, Lynch, A.M.3, and Jeff Hardin3,4
1Department of Biological Sciences, University of Mary Washington, 1301 College Ave., Fredericksburg, VA 22401, USA, 2Department of Biology, SUNY Geneseo, 1 College Cir., Geneseo, NY 14454, USA, 3Program in Genetics, University of Wisconsin, 1117 West Johnson Street, Madison, Wisconsin 53706, 4Department of Zoology, University of Wisconsin, 1117 West Johnson Street, Madison, Wisconsin 53706
Supplemental material for the following paper:
Grana et al (2010). SAX-7/L1CAM and HMR-1/cadherin function redundantly in blastomere compaction and non-muscle myosin accumulation during Caenorhabditis elegans gastrulation. Dev. Biol. 344:731–744. PubMed
Summary
Supplemental Figure 1
Supplemental Figure 2
Supplemental Figure 3
Supplemental Figure 4
Supplemental Figure 5
Supplemental Tables
Movie 1
Movie 2
Movie 3
Movie 4
Movie 5
Movie 6
Summary
Gastrulation is the first major morphogenetic movement in development and requires dynamic regulation of cell adhesion and the cytoskeleton. Caenorhabditis elegans gastrulation begins with the migration of the two endodermal precursors, Ea and Ep, from the surface of the embryo into the interior. Ea/Ep migration provides a relatively simple system to examine the intersection of cell adhesion, cell signaling, and cell movement. Ea/Ep ingression depends on correct cell fate specification and polarization, apical myosin accumulation, and Wnt activated actomyosin contraction that drives apical constriction and ingression (Lee et al., 2006; Nance et al., 2005). Here, we show that Ea/Ep ingression also requires the function of either HMR-1/cadherin or SAX-7/L1CAM. Both cadherin complex components and L1CAM are localized at all sites of cell-cell contact during gastrulation. Either system is sufficient for Ea/Ep ingression, but loss of both together leads to a failure of apical constriction and ingression. Similar results are seen with isolated blastomeres. Ea/Ep are properly specified and appear to display correct apical-basal polarity in sax-7(eq1);hmr-1(RNAi) embryos. Significantly, in sax-7(eq1);hmr-1(RNAi) embryos, Ea and Ep fail to accumulate myosin (NMY-2::GFP) at their apical surfaces, but in either sax-7(eq1) or hmr-1(RNAi) embryos, apical myosin accumulation is comparable to wild type. Thus, the cadherin and L1CAM adhesion systems are redundantly required for localized myosin accumulation and hence for actomyosin contractility during gastrulation. We also show that sax-7 and hmr-1 function are redundantly required for Wnt-dependent spindle polarization during division of the ABar blastomere, indicating that these cell surface proteins redundantly regulate multiple developmental events in early embryos.
Return to top

Supplementary Figure 1. Contact length and
circularity measurements on blastomeres.
Supplemental Figure 1 - PDF (61 Kb) Download
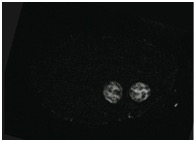
Supplementary Figure 2. END-3::GFP, a marker of gut differentiation, is expressed in sax-7(eq1);hmr-1(RNAi) embryos. Scale bar = 10 μm.
Supplemental Figure 2 - PDF (225 Kb) Download
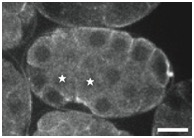
Supplementary Figure 3. PAR localization indicates Ea/Ep are correctly polarized in sax-7(eq1);hmr-1(RNAi) embryos.
PAR-6::GFP was imaged in Ea/Ep shortly following P4 birth in wild-type and mutant embryos. Although GFP distribution was monitored throughout all focal planes, single focal planes are shown. The stars indicate the cells Ea/Ep. Note that in this figure, anterior is to the right. Scale bar = 10 μm.
Supplemental Figure 3 - PDF (315 Kb) Download
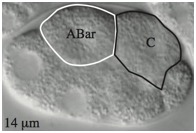
Supplementary Figure 4. Weakened adhesion leads to delayed contact between C and ABar and failure to orient the ABar division.
(A) By 2 min after the birth of C in wild-type embryos, C extends and touches ABar. The surface area of contact rapidly increases, reorienting the ABar spindle. ABar begins to divide 5–6 min after C birth. Correct ABar division orientation is perpendicular to that of ABpr, as shown at time 7 in wild type. When ABar fails to be oriented by a Wnt signal, it divides along an axis nearly parallel to ABpr. This defect in division orientation occurs often in hmr-1(RNAi) embryos and even more frequently in sax-7(eq1);hmr-1(RNAi) embryos. Depth of images is shown at lower left. (B) Time of C contact with ABar with respect to ABar nuclear breakdown. n values are shown. Wild-type ABar orientation is shown as solid circles and abnormal ABar division orientation is shown as open circles. C contact with ABar is delayed in hmr-1(RNAi) embryos, resulting in some abnormal divisions. This delayed contact is even more pronounced in sax-7(eq1);hmr-1(RNAi) embryos (double mutant significantly different from hmr-1(RNAi) alone, p < 0.05). (Embryos were only analyzed for contact when C was clearly visible; thus total numbers here differ from Table S2.)
Supplemental Figure 4 - PDF (2.2 Mb) Download
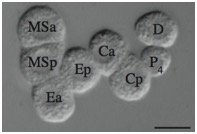
Supplementary Figure 5. Randomization of P2 division orientation in sax-7(eq1);hmr-1(RNAi) blastomeres. As shown in the first panel, in wild-type blastomeres P3 is always anterior to C, while in sax-7(eq1);hmr-1(RNAi), P3 is posterior to C one half of the time. As shown in the second panel, when P3 divides, P4 is anterior to D in both wild-type and sax-7(eq1);hmr-1(RNAi) blastomeres.
Supplemental Figure 5 - PDF (0.9 Mb) Download
Supplemental Tables
Table 1. Timing of Ea/Ep division following loss of function of the cadherin–catenin complex and SAX-7/L1 CAM.
Table 2. Synergy between hmr-1(RNAi) and potential L1CAM linker proteins.
Table 3. ABar spindle defects following CCC and SAX-7/L1CAM loss of function.
Supplemental Tables - PDF (54 Kb) Download

Movie S1 (1 Mb). Nomarski time-lapse movie of a wild-type embryo during Ea/Ep ingression.
Download

Movie S2 (1.1 Mb). Nomarski time-lapse movie of Ea and Ep in a sax-7(eq1); hmr-1(RNAi) embryo at a comparable time to Movie 1.
Download
Return to top
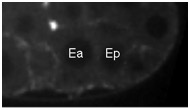

Movie S3 (229 Kb). Confocal time-lapse movie of NMY-2::GFP in a wild-type embryo beginning just before Ea/Ep ingression.
Supplemental NMY-2 moves demonstrate NMY-2 levels over a 15 minute interval between P4 birth and the time of Ea/Ep ingression. Movies were synchronously staged by placing MSxx nuclear reformation at time eight. ImageJ plugins generated by T. Collins at the McMaster Biophotonics Facility were used to correct for photobleaching (available at http://www.macbiophotonics.ca/).
Download
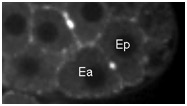
Movie S4 (274 Kb). Confocal time-lapse movie of NMY-2::GFP in a hmr-1(RNAi) embryo beginning just before Ea/Ep ingression.
Download
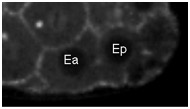
Movie S5 (262 Kb). Confocal time-lapse movie of NMY-2::GFP in a sax-7(eq1) embryo beginning just before Ea/Ep ingression.
Download
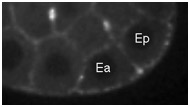
Movie S6 (254 Kb). Confocal time-lapse movie of NMY-2::GFP in a sax-7(eq1); hmr-1(RNAi) embryo at a time comparable to Movies 3, 4, and 5.
Download
Return to top
 THE HARDIN LAB
THE HARDIN LAB